Nouvelles publications
All ===> 10
-
15/10/2020
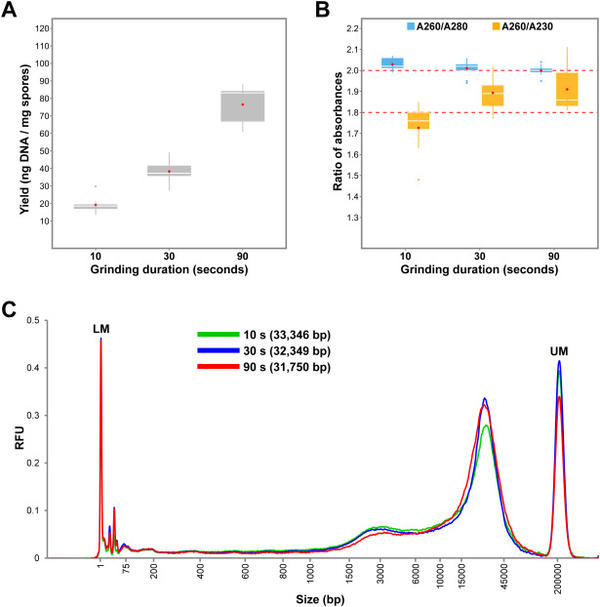
Penouilh-Suzette, C., Fourre, S., Besnard, G., Godiard, L., Pecrix, Y. 2020. A simple method for high molecular-weight genomic DNA extraction suitable for long-read sequencing from spores of an obligate biotroph oomycete. Journal of Microbiological Methods, 178: 106054 https://doi.org/10.1016/j.mimet.2020.106054
-

13/10/2020
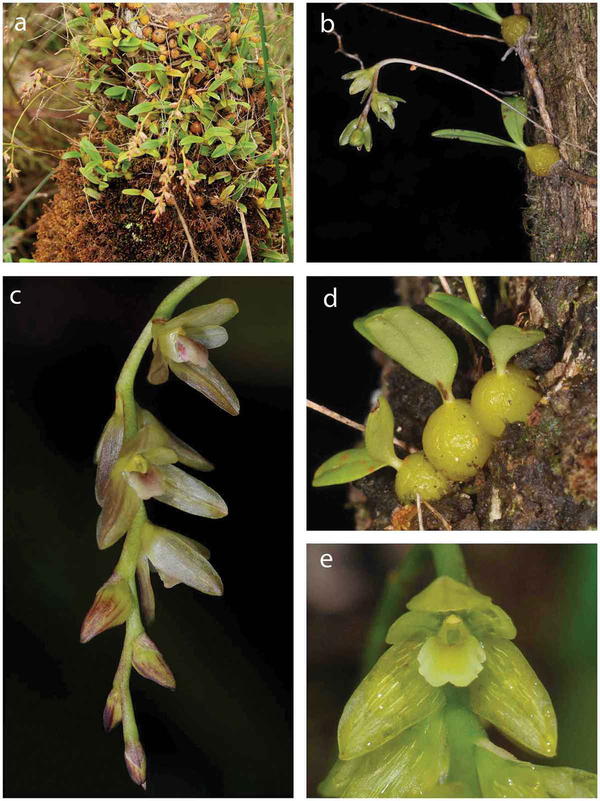
Pailler T., Baider C. 2020: Bulbophyllum mascarenense Pailler and Baider sp nov.: a new endemic orchid species from the Mascarenes, Botany Letters, DOI: 10.1080/23818107.2020.1817145
-
12/10/2020
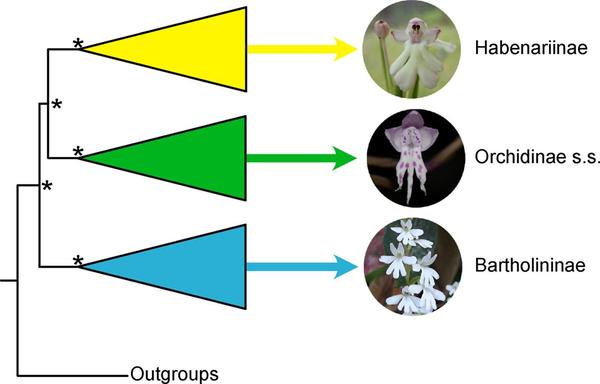
Ngugi G., Le Péchon T., Martos F., Pailler T., Bellstedt D.U., Byterbier B. 2020. Phylogenetic relationships amongst the African genera of subtribe Orchidinae s.l. (Orchidaceae; Orchideae): Implications for subtribal and generic delimitations. Molecular phylogenetics and evolution, 153 : 1-8. https://doi.org/10.1016/j.ympev.2020.106946
-
07/10/2020
Robène, I., Maillot-Lebon, V., Chabirand, A. , Moreau A, Becker, N., Moumène A., Rieux A., Campos P., Gagnevin L., Gaudeul M., Baider C., Chiroleu F., Pruvost O.2020. Development and comparative validation of genomic-driven PCR-based assays to detect Xanthomonas citri pv. citri in citrus plants. BMC Microbiology 20, 296. https://doi.org/10.1186/s12866-020-01972-8
-
06/10/2020
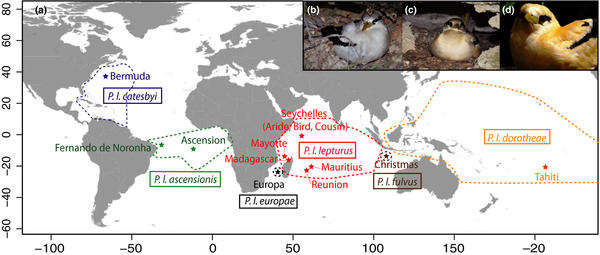
Humeau L., Le Corre M., Reynolds S.J, Wearn C., Hennicke J.C., Russell J.C., Gomard Y., Magalon H. ,Pinet P., Gélin P. , Couzi F.X., Bemanaja E., Tatayah V., Ousseni B., Rocamora G., Talbot P., Shah N., Bugoni L., Da Silva D., Jaeger A. 2020.Genetic structuring among colonies of a pantropical seabird: Implication for subspecies validation and conservation, Ecology and evolution : 1-20 (early view) https://doi.org/10.1002/ece3.6635
-
05/10/2020
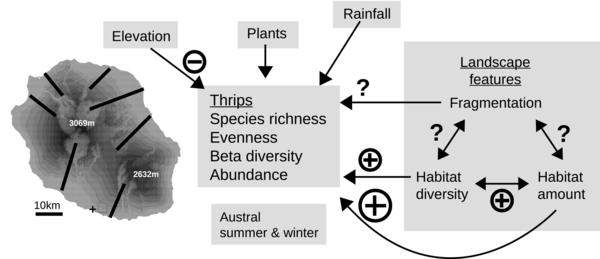
Dianzinga N.T., Moutoussamy M.‐L., Sadeyen J., Ravaomanarivo L.H.R., Frago E. 2020.The interacting effect of habitat amount, habitat diversity and fragmentation on insect diversity along elevational gradients. J Biogeography: 1– 15. https://doi.org/10.1111/jbi.13959
-
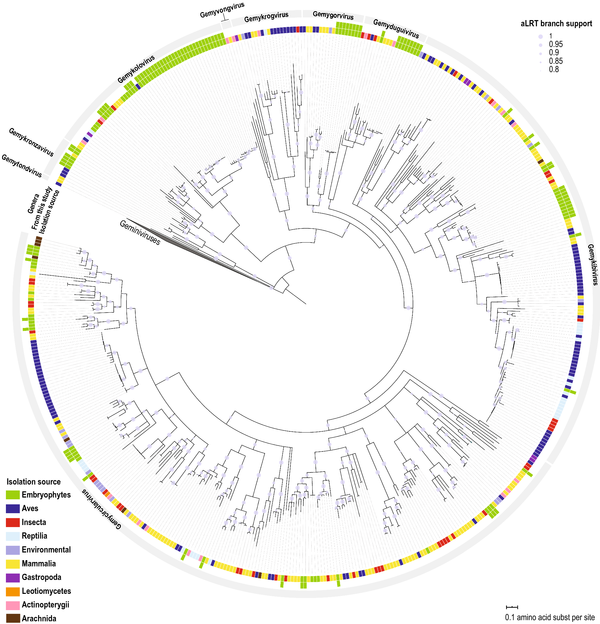
Diverse genomoviruses representing twenty-nine species identified associated with plants
14/09/2020
Fontenele, R.S., Roumagnac, P., Richet, C. , Kraberger S., Stainton D., Aleamotu‘a M., Filloux D., Bernardo P., Harkins G.W, Mc Carthy J., Charles L.S., Lamas N., Abreu E.F.M., Abreu R.A, Batista G, Lacerda A.L.M., And Salywon A., Wojciechowski M.F., Majure L.C., Martin D.P., Ribeiro S.G., Lefeuvre P., Varsani A. 2020 .Diverse genomoviruses representing twenty-nine species identified associated with plants. Arch Virol (https://doi.org/10.1007/s00705-020-04801-5
-

08/09/2020
Fenouillas P., Ah-Peng C., Amy E., Bracco I., Gosset M., Ingrassia F., Lavergne C., Lequette B., Notter J.C., Pausé J.M., Payet N., Payet G., Picot F., Poungavanon N., Strasberg D., Thomas H., Triolo J., Turquet V., Rouget M. (2020). Priorisation spatiale des actions de gestion des plantes exotiques envahissantes: une étape-clé de la conservation à long terme des milieux naturels à La Réunion. CIRAD, Saint Pierre
-
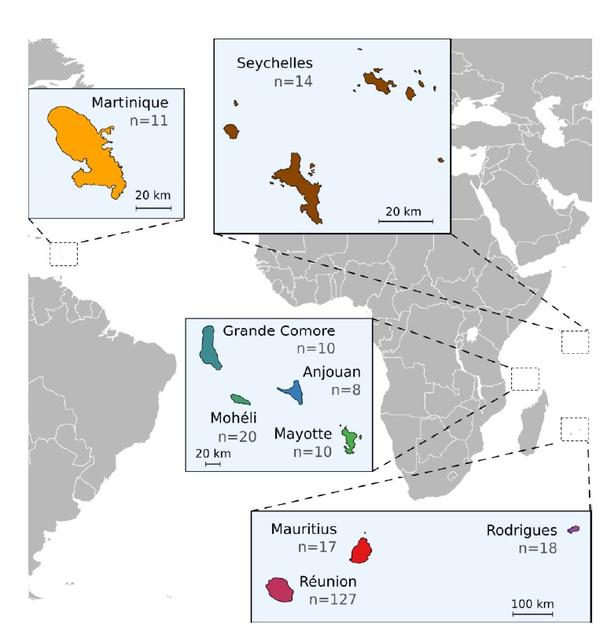
08/09/2020
Richard D., Pruvost O., Balloux F., Boyer C., Rieux A., Lefeuvre P. 2020.Time-calibrated genomic evolution of a monomorphic bacterium during its establishment as an endemic crop pathogen , bioRxiv https://doi.org/10.1101/2020.05.26.115717 (article en preprint)
-
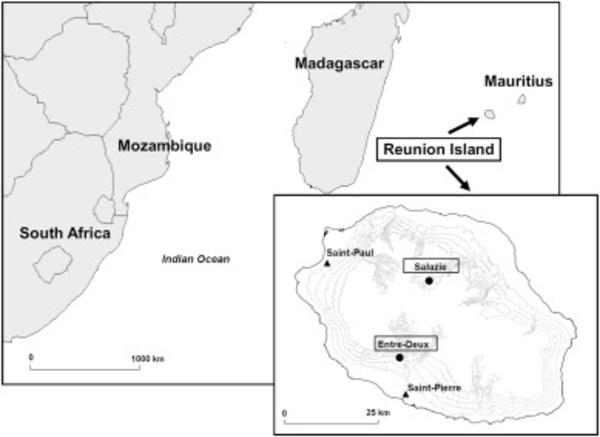
Qualitative modeling of fruit fly injuries on chayote in Réunion: Development and transfer to users
31/08/2020
Deguine, J.-P., Robin, M.H. Corrales, D.C., Vedy-Zecchini, M.-A.,Doizy, A., Chiroleu, F., Quesnel, G., Païtard, I., Bohanec, M., Aubertot, J.-N.2020 Qualitative modelingof fruit fly injuries on chayote in Réunion: Development and transfer to users, Crop Protection (2020),doi: https://doi.org/10.1016/j.cropro.2020.105367. (in press, pre-proof)
Nombre de pages : 46
Nombre éléments : 456
Page courante : 18
- Première page
- Page précédente
- 1
- ...
- 16
- 17
- 18
- ...
- 46
- Page suivante
- Dernière page
